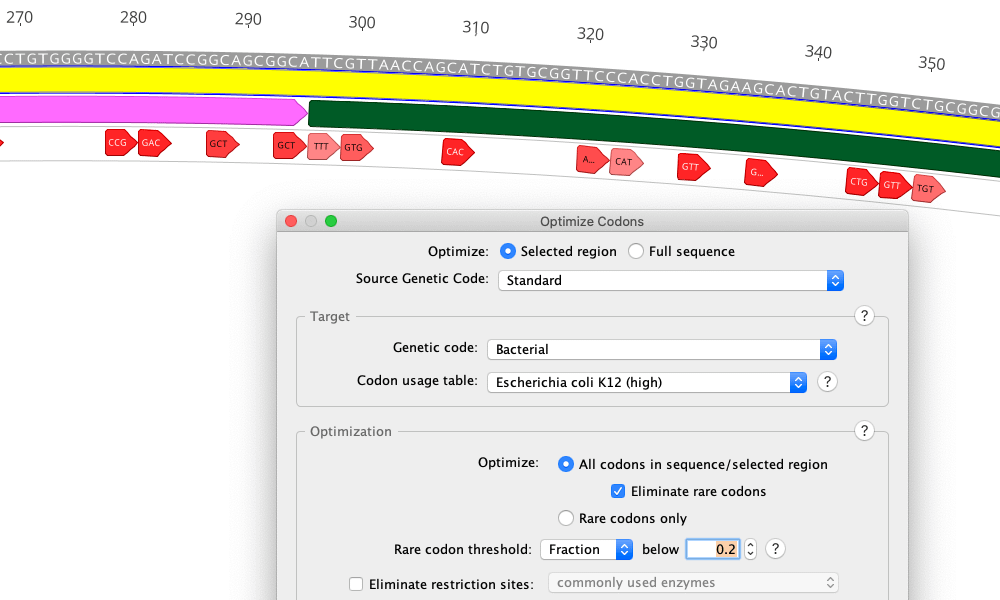
Comprehensive biochemistry analysis tool with advanced molecular cloning Also, thanks to the PlasMapper you can automatically annotate plasmid maps and expression vectors. Moreover, Geneious Prime can help you design primers, simulate PCR and test cloning strategies using smart tools for Gibson and restriction cloning. You can even read arbitrarily large documents like NGS reads or contig assemblies. Hence, you can assemble multiple combinations of Sanger, 454, SOLiD and Illumina data at the same time. Furthermore, Geneious Prime is capable to handle data from various types of sequencing machines with reads of different lengths, including paired-reads and mixed reads from different sequencing machines. The versatile Geneious Assemble is designed to manage read errors caused by incorrect bases or short indels. User-oriented graphical interface and trustworthy reference mapping

On top of that, you can enjoy the eye-catching genome browser, sequence assembly and reference mapping. Regardless of your data type, you can use Geneious Prime to quickly assemble it using fine-tuned and advanced algorithms. What is more, you can display phylogeny along with the alignment and link back to a tree, and use the highlighting feature to quickly locate disagreements. You can also view annotations on the alignments, remove or insert gaps, strip columns, fix bad calls and join sequences. Thanks to the visual sequence alignment and editing you can effortlessly choose the desired algorithms to precisely align sequences without working with multiple files.

Intuitive and easy-to-use sequence editing and analysis tools Most notably, the latest releases of Java do not support 32-bit systems and as a result we currently have to bundle Geneious 32-bit with an old version of Java that is known to have bugs that are addressed in later releases.Geneious Prime is a well-designed and powerful sequence alignment, assembly and analysis application for genomics, microbiology, virology, infectious disease, synthetic biology, ecology, conservation genetics, population genetics, evolutionary biology, agriculture, as well as crop and plant science. Maintaining 32-bit support has become difficult because of the increasingly restricted support that is available for 3rd party tools that Geneious relies on.
Geneious prime 2020 download#
Where can I download previous versions of Geneious Prime for 32-bit?įrom our Previous Versions page.
Please refer to this Microsoft article: 32-bit and 64-bit Windows: Frequently asked questions.
Geneious prime 2020 upgrade#
Am I running 32-bit Windows and how do I upgrade if so? If you are not sure what version of Geneious Prime you are using, check under Help -> About Geneious Prime. This means any users on 64-bit Windows will have to run 64-bit Geneious Prime. People who are unable to upgrade at this time should continue using Geneious Prime 2019.Īdditionally, installers for the 32-bit version of Geneious Prime will no longer be available for version 2020 onwards. To continue using the latest version of Geneious Prime, users of 32-bit Windows will need to upgrade to a 64-bit operating system. Geneious Prime will no longer run on 32-bit Windows operating systems from version 2020 onwards (scheduled for release late 2019).

